ECG Analysis Using Deep CNNs
Introduction
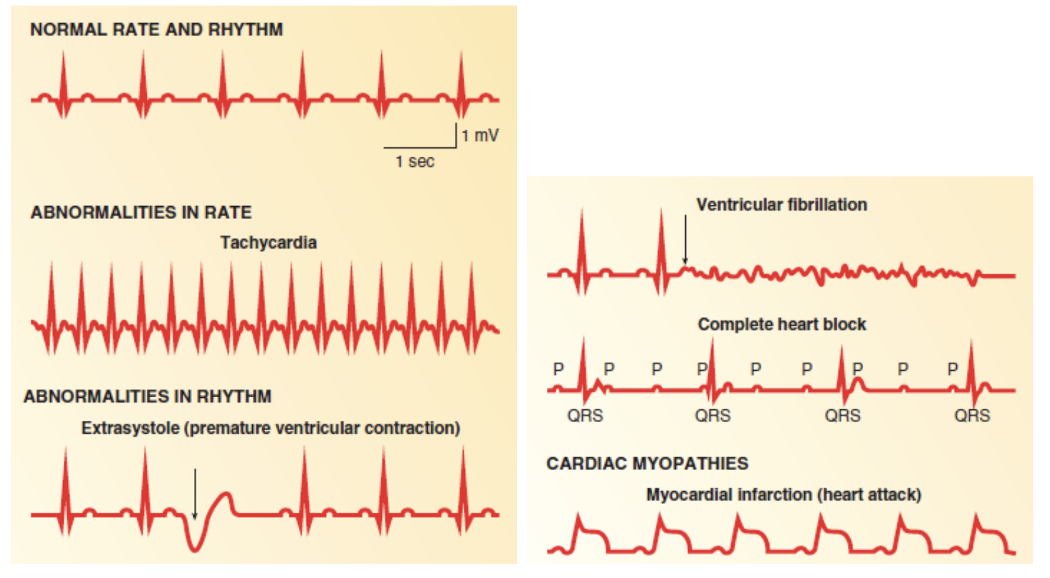
Cardiovascular disease (CVD) is the leading cause of death in the world, representing 32% of all global deaths. One of the most common abnormalities associated with CVDs are heart arrhythmias, diagnosed using electrocardiogram (ECG) recordings. In this project, Yan Zhu and her team used machine learning to classify ECG heartbeats by testing two different CNN-based approaches and compare the two methods. ECGs are helpful to doctors in diagnosing arrhythmias non-invasively and deciding on the best course of treatment. However, distinguishing heartbeats on ECGs is time-consuming and difficult, as they usually contains lots of noise and are prone to misinterpretation. By using a machine learning model to aid in this task, it will help with the speed and accuracy of diagnoses.
Framework
Dataset
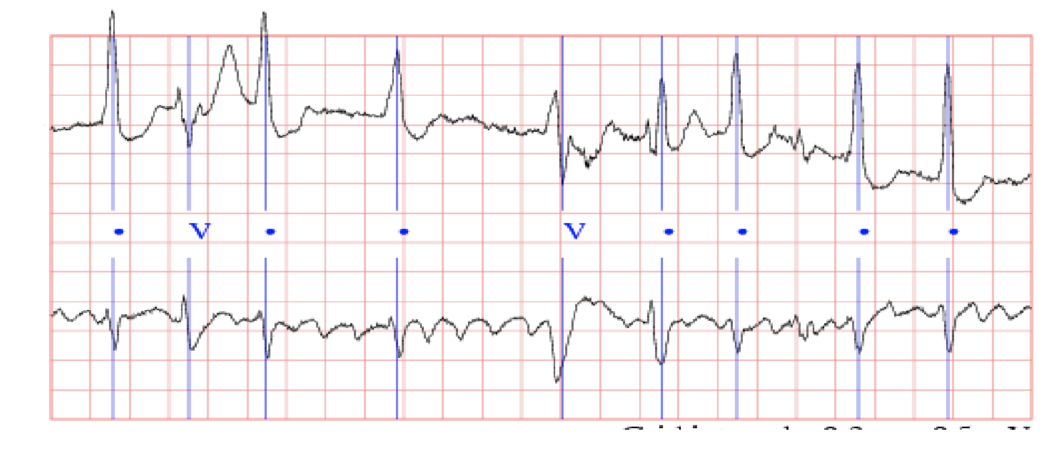
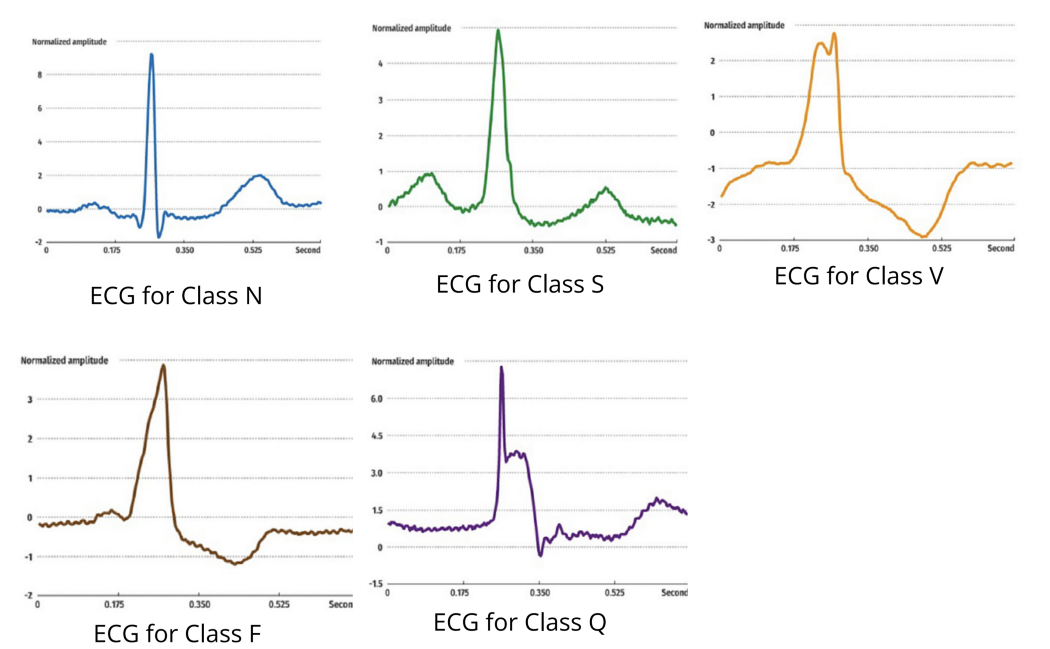
Yan and her team used the PhysioBank MIT-BIG arrhythmia database, containing 48 half-hour excerpts of ECG recordings from 47 subjects. These subjects were studied by the BIH Arrhythmia Laboratory between 1975 and 1979, and the dataset has been widely used in many biomedical/machine learning projects.
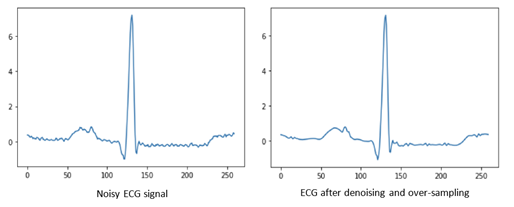
Preprocessing
The resulting data was preprocessed in two steps. In the first step, a wavelet transform was used to denoise the signal. In the second step, the Pan-Tompkins algorithm was used to detect R peaks and segment the ECG signal into heartbeats. The wavelet transform then utilized a daubechies-6 wavelet decomposition transform. The Daubechies wavelets are a family of orthogonal wavelets defining a discrete wavelet transform and characterized by a maximal number of vanishing moments for some given support. The resulting wavelets, after the transform, are noise-free and retain the necessary diagnostics information contained in the original ECG signal, and therefore, can deliver the heartbeat information more precisely.
Architecture
Model 1 - CNN
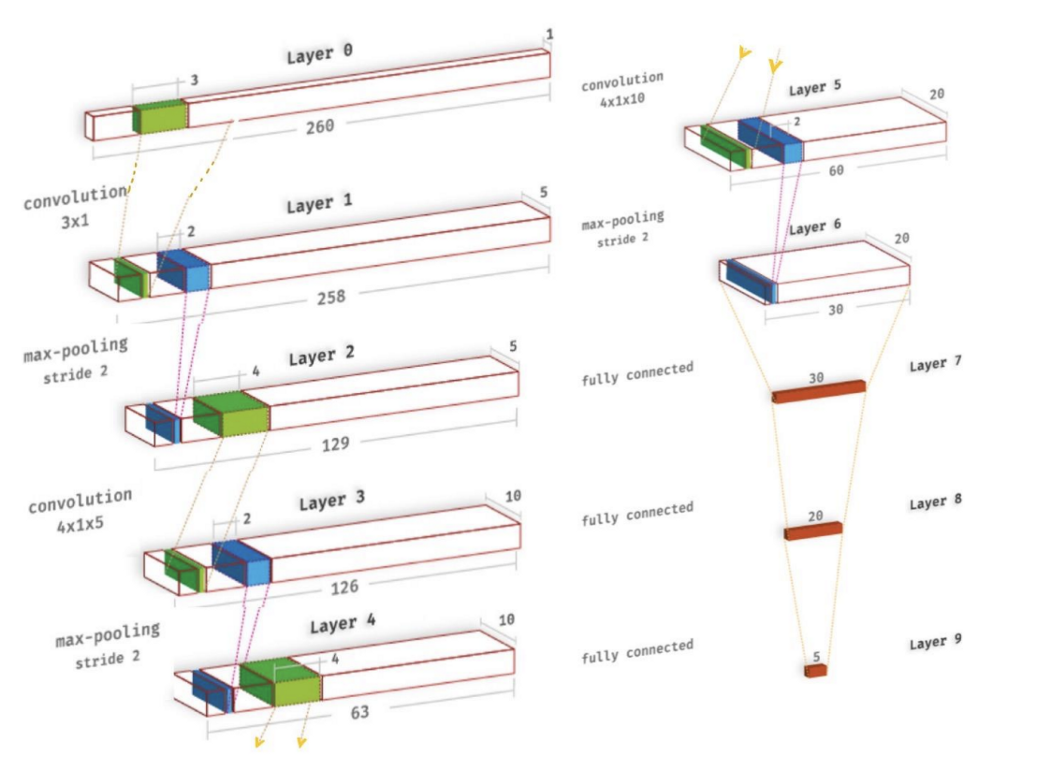
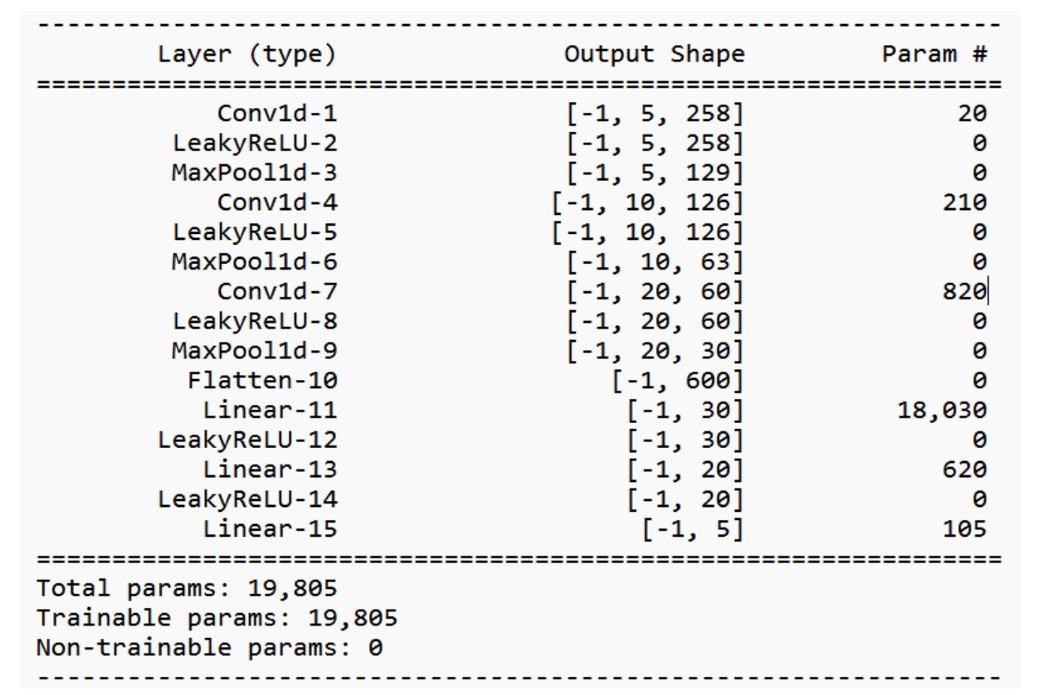
The convolutional neural network implemented was based on the one by Acharya et al. There are three one-dimensional convolutional layers, each of which uses a Leaky-ReLU activation and is followed by a max-pooling layer. The convolutional layers are important for picking up the key features of the heartbeat sequences, such as the QRS complexes, as well as the locations of different kinds of arrhythmia. Afterwards, the output feature map gets passed into a 3-layer fully-connected network, also using Leaky-ReLUs, followed by a softmax in the final layer.
Model 2 - CNN-LSTM Combination
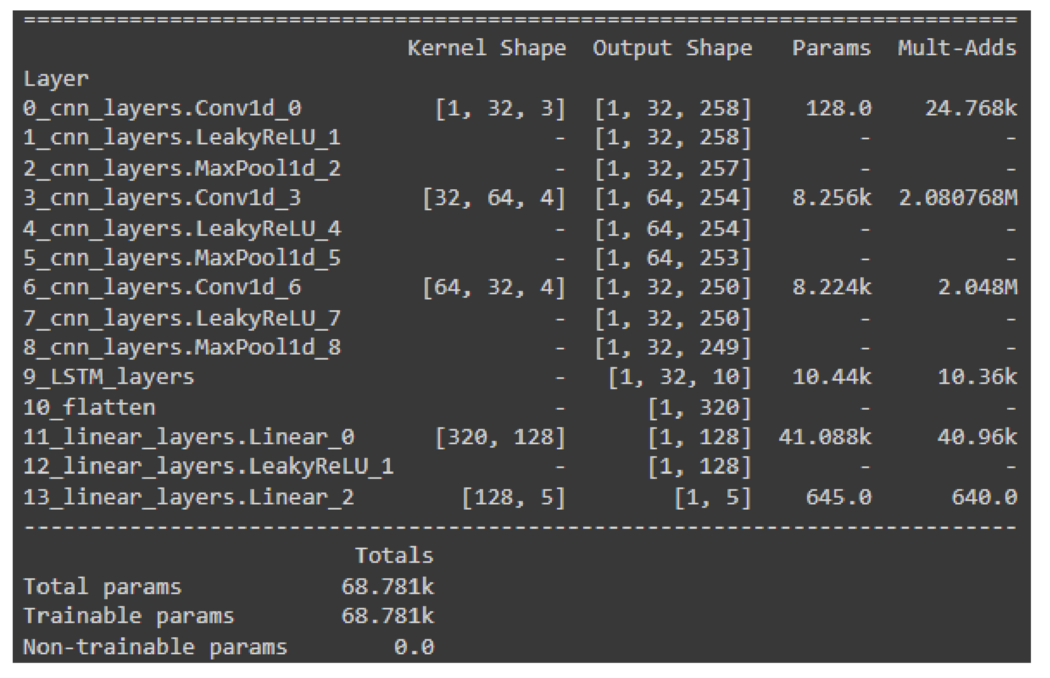
In CNNs, long-distance relations are lost due to the local nature of the convolution kernels. For that reason, when dealing with sequential data, recurrent neural networks (RNNs) models are preferred over CNNs. To leverage the inductive biases of both architectures, in the LSTM model, firstly, convolutional layers are applied to the data for denoising and local feature extraction purposes. Then, the extracted features are fed into LSTM layers so that the sequential nature of the data is exploited. The LSTM model comprises three initial 1D convolutional layers followed by the LSTM layer and two dense linear layers. For all layers, the leaky ReLU activation function is employed.
Future Works
We are continuiing this project in 2022 as “12-Lead ECG Reconstruction”, where we aim to reconstruct the full ECG signal (captured with 12 leads) from only 2-3 leads. We will be collaborating with ONHand AI for additional dataset with greature accuracy and resolution. For more details, please visit https://utmist.gitlab.io/projects/12-lead-ecg/.